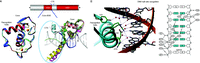Understanding nuclear receptor form and function using structural biology
-
Figure 1
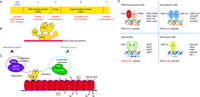
Domain organization of NRs and their interaction partners. (A) NR polypeptides have an N-terminal domain (NTD) that is variable in size and sequence, a conserved DBD, a variable hinge region, and a 12-helical LBD. Several NRs also contain a variable F-domain positioned at their C-termini. (B) Schematic showing how a dimeric NR uses different domains to bind to DNA and ligand. (C) NRs can be modulated by ligands that either activate or repress gene targets. Repression is mediated by complexes that have corepressors (SMRT/N-CoR and histone deacetylases (HDACs)), among other components. Activation requires the coactivator complexes (p160 family members and histone acetyltransferase (HAT)). The repressive and activating complexes block or promote transcription. (D) Oligomeric complexes of NRs and DNA response element repertoires. Receptors can be organized into distinct oligomeric states such as heterodimers with the common partner retinoid X receptor (RXR), homodimers, or monomers. The non-steroid receptor heterodimers and many homodimers bind to direct repeat response elements with various inter-half-site spacings. Steroid receptor homodimers mainly use palindromic DNA elements, where the two half sites are in an inverted repeat arrangement. Other receptors use monomeric sites extended at their 5′-end with short sequences used for selectivity. Several examples of receptors falling into each of these four categories are shown.
-
Figure 2
Understanding domain–domain integration in nuclear receptors requires structural studies that utilize the complete receptor complexes. A major goal has been to understand allosteric communication: how signals in one domain may be efficiently transmitted to a distal domain in the quaternary fold (Chandra et al. 2008, 2013).
-
Figure 3
The basis for DNA recognition. (A) The core DBD is the 66-residue region with two zinc-binding modules. This domain is responsible for the recognition of hexameric DNA half sites. Shown on the left are the core DBD structures of RXR (blue) and RAR (red) superimposed. The C-terminal extension (CTE) of the DBD, which lies within the immediate hinge region, can also participate in DNA binding and spacer recognition. Shown on the right is the superposition of the DBD-CTE segments of NGFI-B (green), LRH-1 (cyan), VDR (yellow), TR (magenta), and Rev-Erb (salmon). While the core DBDs are well conserved in folding, the CTE sequences and their structures are divergent. The CTEs can act as discriminators of DNA spacing. (B) RAR DBD interactions with AGGTCA elements. DNA half-site recognition by NRs involves a series of hydrophilic residues positioned on the same face of the DBD recognition helix. These residues read the DNA base pair sequence at the major groove. Other basic amino acids additionally stabilize DNA binding by interacting with the phosphate backbone of the DNA. Crystallographic studies have shown that water molecules (shown as red or black circles) often help mediate DNA contacts.
-
Figure 4
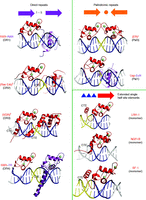
The interactions of DBDs with response elements. Shown are structures of NR DBD homodimers, heterodimers, and monomers on response elements consisting of direct repeats (DRs), palindromes (Pal), and single half sites. The spacer size has a dramatic effect on the relative rotation and displacement of the two DBDs, fostering their productive dimerization contacts on the correctly spaced element, but blocking their dimerization when the spacing size is incorrect. Monomeric NRs use the same consensus half sites but rely on the immediate flanking base pairs upstream of their half site for response element discrimination.
-
Figure 5
Structural properties of NR LBDs. (A) The comparison of the unligand RXRα with the liganded RARγ first suggested that ligands can induce multiple changes in the LBD conformation, most strikingly in the positioning of helix-12 (H12). The movement of H12 (shown in red) allows the ligand-binding pocket to become enclosed to prevent ligand escape, in what can be described as a trapping mechanism. Other smaller rearrangements are additionally induced by ligand binding (blue arrows). (B) Typical dimerization surfaces that form between receptor LBDs. Shown here are the LXR–RXR LBD heterodimer and the ER LBD homodimer. For two LBDs to dimerize, surfaces from H7, H9, and H10/11 participate in forming the dimeric interfaces.
-
Figure 6
Coactivator binding to the surface of the LBD. (A) Shown is the RORβ LBD structure with an agonist ligand (blue) that places helix-12 (H12) in the active conformation. From this position, H12 fosters the interactions of an LXXLL motif contained in most p160 coactivators with the surface of the LBD. The LXXLL motif forms a small helical segment, allowing the leucines along one face to firmly dig into a hydrophobic groove on the surface of the LBD. The X residues in the coactivator motif are typically polar and interact with the solvent. Notice that the ligand (blue) is shielded from solvent when H12 is in the active conformation.
-
Figure 7
Structural understanding of how agonists and antagonists mediate their effects on the estrogen receptor. (A) Both estradiol and the synthetic molecule diethylstilbestrol (DES) are agonists, whereas tamoxifen and raloxifene are antagonists. The antagonists differ from stilbesterol only in the addition of the extension moiety (in blue). (B) DES binding places helix-12 (H12) in the agonist conformation, as does estradiol. (C) The extension moiety in raloxifene pushes H12 out of its active conformation.
-
Figure 8
Comparison of coactivator and corepressor binding modes. (A) The synthetic ligand GW731 is an agonist of PPARα, producing the active conformation and allowing for favorable coactivator binding. (B) In contrast, the synthetic molecule GW6471 displaces helix-12 in this receptor and helps create a stable conformation that allows for a region of the corepressor SMRT to bind onto the surface. (C) The binding of the Rev-Erbα LBD to a corepressor element in the absence of ligand.
-
Figure 9
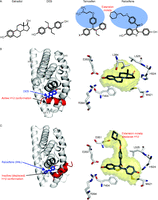
Demonstration of the distinct orientations of cholesterol-derived ligands in receptor pockets. (A) FXR binding to the bile acid CDCA. (B) Details of interactions with solid red lines indicating hydrogen bonds and dotted lines indicating van der Waals interactions. Note that the steroidal A-ring is positioned at helix-12 (H12). (C and D) LXRα binding to epoxycholesterol requires the steroidal A-ring to be in the opposite direction, while its side-chain points at H12.
-
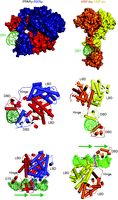
Figure 11
Crystal structures of multi domain NR complexes. On the left side is the PPARγ–RXRα complex on DR1 and on the right side is the HNF-4α homodimer on DR1. RXR is indicated in blue and PPAR is indicated in red. The upstream subunit of HNF-4α is indicated in yellow and the downstream subunit is indicated in orange. The green arrows show the direction of the AGGTCA direct repeats in the DR1 response elements in the lower panel for both complexes. Three views of each complex are shown.
- © 2013 Society for Endocrinology