In vitro model systems to study androgen receptor signaling in prostate cancer
-
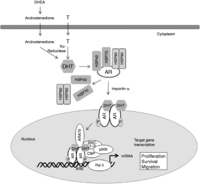
Figure 1
Classic androgen receptor (AR) genomic activity via androgen. Androgens derive predominantly from the testis (90–95%) but also to a lesser extent from the adrenal glands (5–10%) and mediate their effects via binding to the AR. Testicular testosterone (T) and adrenal DHEA or androstenedione are converted locally in the prostate into bioactive DHT by the enzymes 5α-reductase 1 and 2. In the classic mode of AR genomic activity, androgen binding to the AR induces a conformational change that leads to the dissociation of chaperone and heat shock proteins (HSP40, HSP90) and its subsequent interaction with coregulatory molecules and importin-α, which facilitate nuclear translocation of AR–ligand complexes. In the nucleus, the AR undergoes phosphorylation and dimerization, which permits chromatin binding to androgen-responsive elements (ARE) within androgen-regulated target genes. The AR recruits a variety of coactivators (ARA70, SRC-1, -3, and CBP/p300) and RNA polymerase II (Pol II) to induce gene transcription.
-
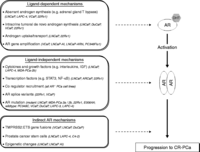
Figure 2
In vitro cell models exhibiting characteristics of androgen receptor (AR)-dependent and AR-independent mechanisms that promote prostate cancer (PCa) progression to castration-resistance. Several pathways have been identified by which PCa cells can overcome androgen depletion and thereby facilitate tumor progression to CR-PCa and can be divided into: i) ligand-dependent mechanisms, which promote AR activation despite castrate levels of androgens; ii) ligand-independent mechanisms, which facilitate AR activation by nonandrogenic factors and/or altering the intrinsic behavior/sensitivity of AR; and iii) indirect mechanisms that act downstream of AR activation (e.g. chromatin remodeling via histone deacetylases, re-emergence of tumors via CSCs and AR-dependent expression of oncogenic ETS transcription factors). Cell lines that have been used to study these different mechanisms are indicated.
- © 2013 Society for Endocrinology